Import Requisite Libraries
######################## Standard Library Imports ############################## import pandas as pdimport osimport sysfrom eda_toolkit import ensure_directory######################## Modeling Library Imports ############################## import shapimport model_tunerfrom model_tuner.pickleObjects import loadObjectsimport eda_toolkitimport matplotlib.pyplot as pltfrom functions import evaluate_kfold_oof, build_multimodel_performance_table# Add the parent directory to sys.path to access 'functions.py' print (f"This project uses: \n \n Python { sys. version. split()[0 ]} \n model_tuner " f" { model_tuner. __version__} \n eda_toolkit { eda_toolkit. __version__} "
/home/lshpaner/Python_Projects/circ_milan/venv_circ_311/lib/python3.11/site-packages/tqdm/auto.py:21: TqdmWarning: IProgress not found. Please update jupyter and ipywidgets. See https://ipywidgets.readthedocs.io/en/stable/user_install.html
from .autonotebook import tqdm as notebook_tqdm
This project uses:
Python 3.11.11
model_tuner 0.0.34b1
eda_toolkit 0.0.19
Set Paths & Read in the Data
# Define base paths # `base_path`` represents the parent directory of current working directory = os.path.join(os.pardir)# Go up one level from 'notebooks' to the parent directory, then into the 'data' folder = os.path.join(os.pardir, "data" )= os.path.join(base_path, "images" , "png_images" , "modeling" )= os.path.join(base_path, "images" , "svg_images" , "modeling" )# Use the function to ensure the 'data' directory exists
Directory exists: ../data
Directory exists: ../images/png_images/modeling
Directory exists: ../images/svg_images/modeling
= "../data/processed/" = "../mlruns/models/"
= pd.read_parquet(os.path.join(data_path, "X.parquet" ))
print (f"DataFrame Columns w/ Outcome: \n { df. columns. to_list()} " )print (f"DataFrame Shape: { df. shape} " )
DataFrame Columns w/ Outcome:
['Age_years', 'BMI', 'Surgical_Technique', 'Intraoperative_Blood_Loss_ml', 'Intraop_Mean_Heart_Rate_bpm', 'Intraop_Mean_Pulse_Ox_Percent', 'Surgical_Time_min', 'BMI_Category_Obese', 'BMI_Category_Overweight', 'BMI_Category_Underweight', 'Intraop_SBP', 'Intraop_DBP', 'Diabetes']
DataFrame Shape: (194, 13)
= pd.read_parquet(os.path.join(data_path, "X.parquet" ))= pd.read_parquet(os.path.join(data_path, "y_Bleeding_Edema_Outcome.parquet" ))= df.join(y, how= "inner" , on= "patient_id" )
Load Models
# lr_smote_training = loadObjects("./192577440515948778/f9e6938832d8403b96c13485638d7ff2/artifacts/lr_Bleeding_Edema_Outcome/model.pkl" ,# rf_over_training = loadObjects("./192577440515948778/fa0e2889515d4f2cb2e37b9a8feef84d/artifacts/rf_Bleeding_Edema_Outcome/model.pkl" ,# svm_orig_training = loadObjects("./192577440515948778/936e228e12834c519a472f4a3556db66/artifacts/svm_Bleeding_Edema_Outcome/model.pkl" ,
Object loaded!
Object loaded!
Object loaded!
Set-up Pipelines, Model Titles, and Thresholds
= [model_lr, model_rf, model_svm]# Model titles = ["Logistic Regression" ,"Random Forest Classifier" ,"Support Vector Machines" ,= {"Logistic Regression" : next (iter (model_lr.threshold.values())),"Random Forest Classifier" : next (iter (model_rf.threshold.values())),"Support Vector Machines" : next (iter (model_svm.threshold.values())),
for col in X.columns:if col.startswith("BMI_" ):print (f"Value Counts for column { col} : \n " )print (X[col].value_counts())print (" \n " )
Value Counts for column BMI_Category_Obese:
BMI_Category_Obese
0 183
1 11
Name: count, dtype: int64
Value Counts for column BMI_Category_Overweight:
BMI_Category_Overweight
0 141
1 53
Name: count, dtype: int64
Value Counts for column BMI_Category_Underweight:
BMI_Category_Underweight
0 190
1 4
Name: count, dtype: int64
SHAP Summary Plot
SHAP (SHapley Additive exPlanations) Set-up
# Step 1: Get transformed features using model's preprocessing pipeline = model_svm.get_preprocessing_and_feature_selection_pipeline().transform(# Optional: Sampling for speed (or just use X_transformed if it's small) = 100 = shap.utils.sample(X_transformed, sample_size, random_state= 42 )# Step 2: Get final fitted model (SVC in pipeline) = model_svm.estimator.named_steps[model_svm.estimator_name]# Step 3: Define a pred. function that returns only the probability for class 1 def model_predict(X):return final_model.predict_proba(X)[:, 1 ]# Step 4: Create SHAP explainer = shap.KernelExplainer(= model_svm.get_feature_names()# Step 5: Compute SHAP values for the full dataset or sample = explainer.shap_values(X_sample) # can use X_transformed instead
100%|██████████| 100/100 [03:17<00:00, 1.97s/it]
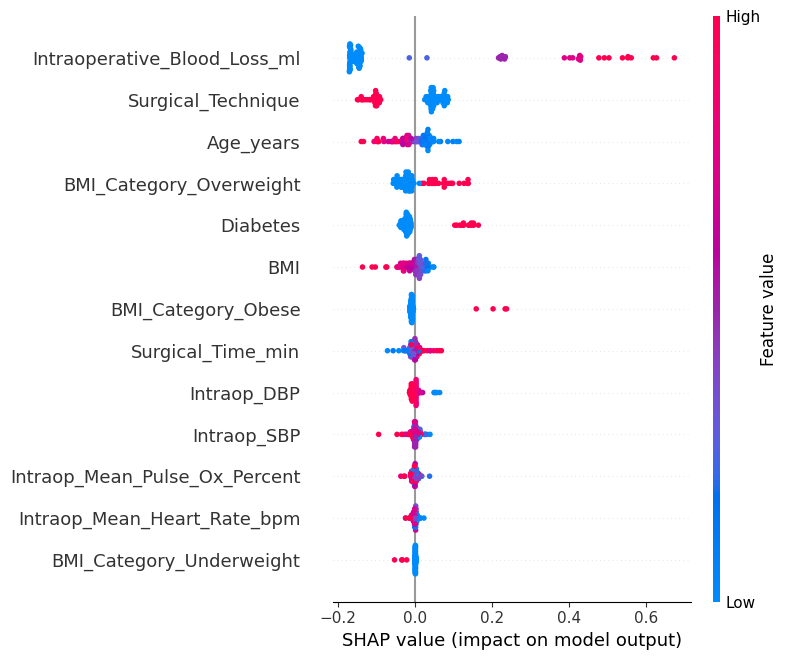
SHAP Beeswarm Plot
# Step 6a: SHAP beeswarm plot (default) = model_svm.get_feature_names(),= False ,"shap_summary_beeswarm.png" ), dpi= 600 )"shap_summary_beeswarm.png" ), dpi= 600 )
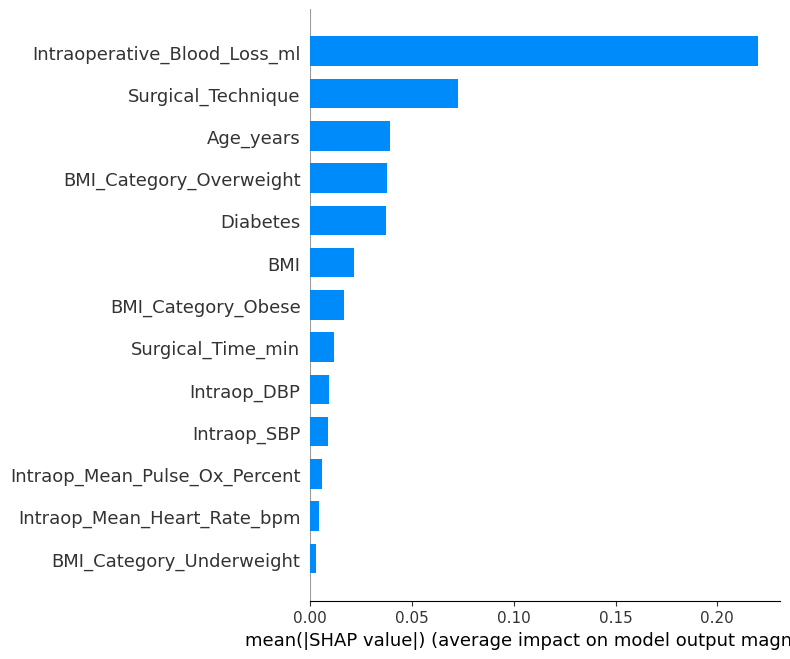
SHAP Bar Plot
# Step 6b: SHAP bar plot (mean |SHAP value| for each feature) = model_svm.get_feature_names(),= "bar" ,= False ,"shap_summary_bar.png" ), dpi= 600 )"shap_summary_bar.png" ), dpi= 600 )
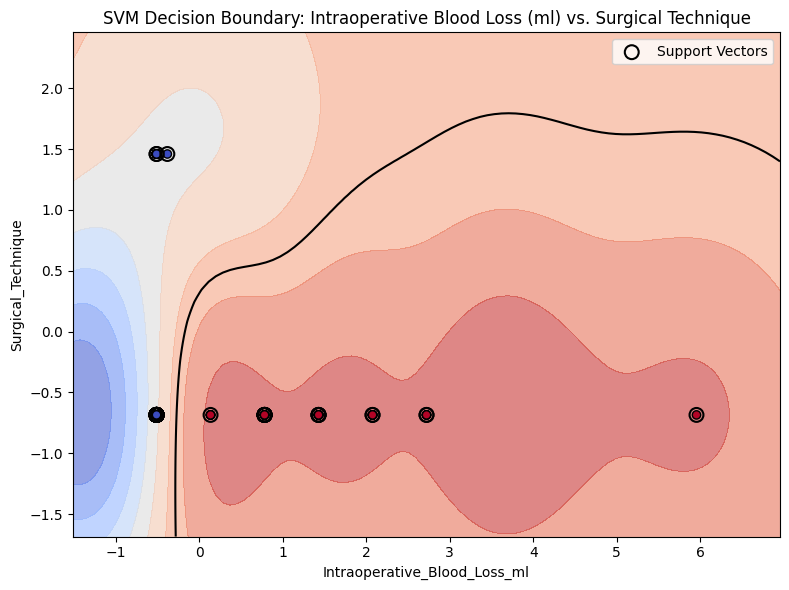
Plot SVM Decision Boundary
from project_functions import plot_svm_decision_boundary_2d# model=model_svm, = X,= y,= ("Intraoperative_Blood_Loss_ml" , "Surgical_Technique" ),= "SVM Decision Boundary: Intraoperative Blood Loss (ml) vs. Surgical Technique" ,= os.path.join(image_path_svg, "svm_decision_surface_2d.svg" ),
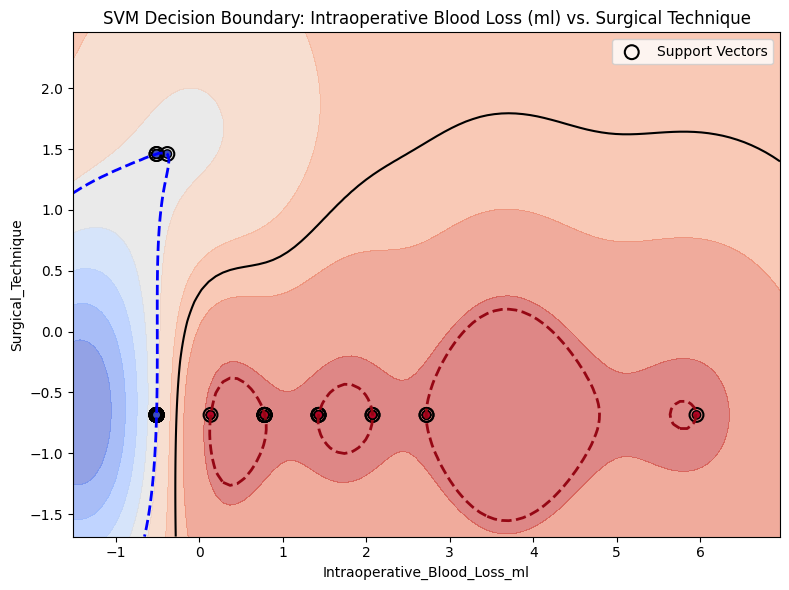
from project_functions import plot_svm_decision_boundary_2d# model=model_svm, = X,= y,= ("Intraoperative_Blood_Loss_ml" , "Surgical_Technique" ),= "SVM Decision Boundary: Intraoperative Blood Loss (ml) vs. Surgical Technique" ,= True ,= os.path.join(image_path_svg, "svm_decision_surface_2d_margin.svg" ),
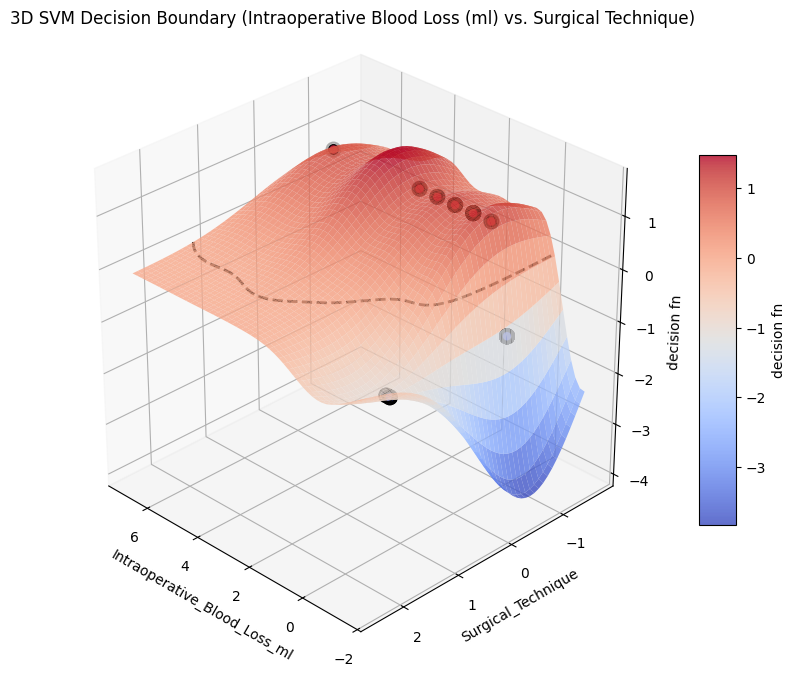
from project_functions import plot_svm_decision_surface_3d= X,= y,# figsize=(6, 10), = ("Intraoperative_Blood_Loss_ml" , "Surgical_Technique" ),= "3D SVM Decision Boundary (Intraoperative Blood Loss (ml) vs. Surgical Technique)" ,= os.path.join(image_path_png, "svm_decision_surface_3d.png" ),= os.path.join(image_path_svg, "svm_decision_surface_3d.svg" ),
from project_functions import plot_svm_decision_surface_3d_plotly# Plotly 3D SVM Decision Surface = df,= df["Bleeding_Edema_Outcome" ],= ("Intraoperative_Blood_Loss_ml" , "Surgical_Technique" ),= f"Interactive 3D SVM Decision Boundary:<br>Intraoperative Blood " f"Loss (ml) vs. Surgical Technique" ,= os.path.join(image_path_svg, "svm_decision_surface_3d_plotly.html" ),
Saved interactive plot to ../images/svg_images/modeling/svm_decision_surface_3d_plotly.html
Calibration
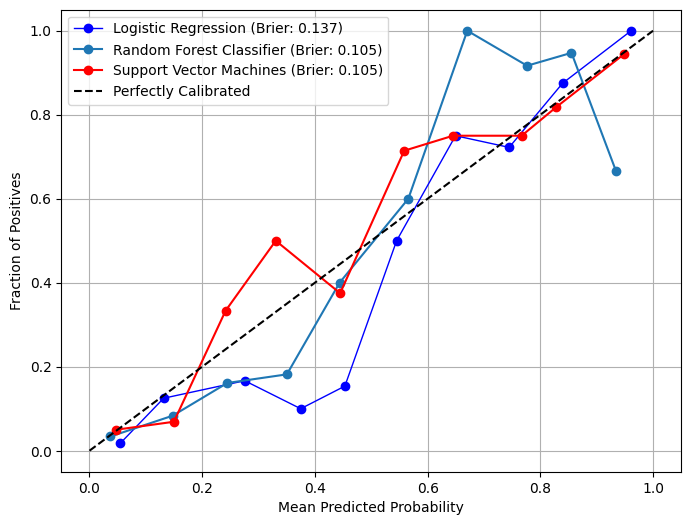
# Plot calibration curves in overlay mode from model_metrics import show_calibration_curve= pipelines_or_models,= X,= y,= model_titles,= True ,= "" ,= True ,= image_path_png,= image_path_svg,= 40 ,= {"Logistic Regression" : {"color" : "blue" , "linewidth" : 1 },"Support Vector Machines" : {"color" : "red" ,# "linestyle": "--", "linewidth" : 1.5 ,"Decision Tree" : {"color" : "lightblue" ,"linestyle" : "--" ,"linewidth" : 1.5 ,= (8 , 6 ),= 10 ,= 10 ,= 10 ,= True ,= 3 ,= False ,# gridlines=False, = {"color" : "black" },
Running k-fold model metrics...
Processing Folds: 100%|██████████| 10/10 [00:02<00:00, 3.42it/s]
Running k-fold model metrics...
Processing Folds: 100%|██████████| 10/10 [00:02<00:00, 3.65it/s]
Running k-fold model metrics...
Processing Folds: 100%|██████████| 10/10 [00:00<00:00, 39.68it/s]
Confusion Matrices
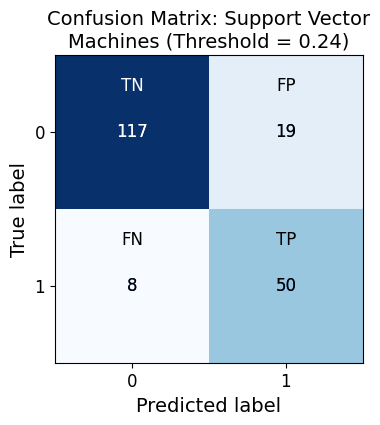
from model_metrics import show_confusion_matrix= pipelines_or_models,= X,= y,= model_titles,= [thresholds],# class_labels=["No Pain", "Class 1"], = "Blues" ,= 40 ,= True ,= image_path_png,= image_path_svg,= True ,= 3 ,= 1 ,= (4 , 4 ),= False ,= 14 ,= 12 ,= 12 ,= True ,# thresholds=thresholds, # custom_threshold=0.5, # labels=False,
Running k-fold model metrics...
Processing Folds: 100%|██████████| 10/10 [00:00<00:00, 30.43it/s]
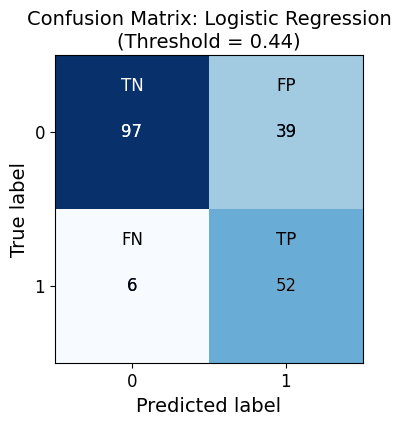
Confusion Matrix for Logistic Regression:
Predicted 0 Predicted 1
Actual 0 97 39
Actual 1 6 52
Classification Report for Logistic Regression:
precision recall f1-score support
0 0.94 0.71 0.81 136
1 0.57 0.90 0.70 58
accuracy 0.77 194
macro avg 0.76 0.80 0.75 194
weighted avg 0.83 0.77 0.78 194
Running k-fold model metrics...
Processing Folds: 100%|██████████| 10/10 [00:02<00:00, 3.74it/s]
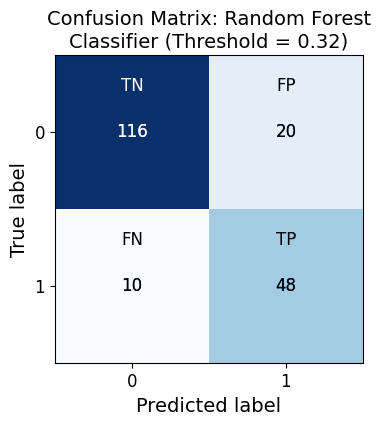
Confusion Matrix for Random Forest Classifier:
Predicted 0 Predicted 1
Actual 0 116 20
Actual 1 10 48
Classification Report for Random Forest Classifier:
precision recall f1-score support
0 0.92 0.85 0.89 136
1 0.71 0.83 0.76 58
accuracy 0.85 194
macro avg 0.81 0.84 0.82 194
weighted avg 0.86 0.85 0.85 194
Running k-fold model metrics...
Processing Folds: 100%|██████████| 10/10 [00:00<00:00, 53.64it/s]
Confusion Matrix for Support Vector Machines:
Predicted 0 Predicted 1
Actual 0 117 19
Actual 1 8 50
Classification Report for Support Vector Machines:
precision recall f1-score support
0 0.94 0.86 0.90 136
1 0.72 0.86 0.79 58
accuracy 0.86 194
macro avg 0.83 0.86 0.84 194
weighted avg 0.87 0.86 0.86 194
ROC AUC Curves
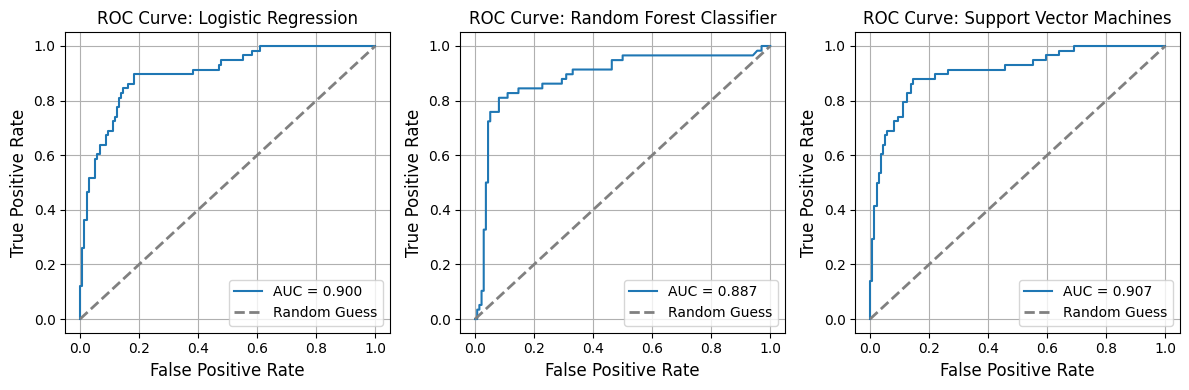
from model_metrics import show_roc_curve# Plot ROC curves = pipelines_or_models,= X,= y,= False ,= model_titles,= 3 ,# n_cols=3, # n_rows=1, # curve_kwgs={ # "Logistic Regression": {"color": "blue", "linewidth": 2}, # "SVM": {"color": "red", "linestyle": "--", "linewidth": 1.5}, # }, # linestyle_kwgs={"color": "grey", "linestyle": "--"}, = True ,= True ,= 3 ,= (12 , 4 ),# label_fontsize=16, # tick_fontsize=16, = image_path_png,= image_path_svg,# gridlines=False,
Running k-fold model metrics...
Processing Folds: 100%|██████████| 10/10 [00:00<00:00, 42.06it/s]
AUC for Logistic Regression: 0.900
Running k-fold model metrics...
Processing Folds: 100%|██████████| 10/10 [00:02<00:00, 4.42it/s]
AUC for Random Forest Classifier: 0.887
Running k-fold model metrics...
Processing Folds: 100%|██████████| 10/10 [00:00<00:00, 63.01it/s]
AUC for Support Vector Machines: 0.907
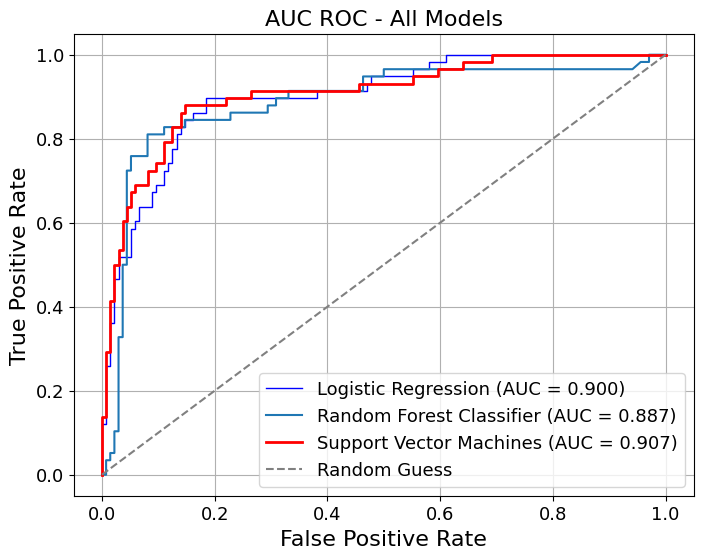
= pipelines_or_models,= X,= y,= True ,= model_titles,= "AUC ROC - All Models" ,= {"Logistic Regression" : {"color" : "blue" , "linewidth" : 1 },"Random Forest" : {"color" : "lightblue" , "linewidth" : 1 },"Support Vector Machines" : {"color" : "red" ,"linestyle" : "-" ,"linewidth" : 2 ,= {"color" : "grey" , "linestyle" : "--" },= True ,= False ,= 3 ,= (8 , 6 ),# gridlines=False, = 16 ,= 13 ,= image_path_png,= image_path_svg,
Running k-fold model metrics...
Processing Folds: 100%|██████████| 10/10 [00:00<00:00, 39.27it/s]
AUC for Logistic Regression: 0.900
Running k-fold model metrics...
Processing Folds: 100%|██████████| 10/10 [00:02<00:00, 4.51it/s]
AUC for Random Forest Classifier: 0.887
Running k-fold model metrics...
Processing Folds: 100%|██████████| 10/10 [00:00<00:00, 48.48it/s]
AUC for Support Vector Machines: 0.907
Precision-Recall Curves
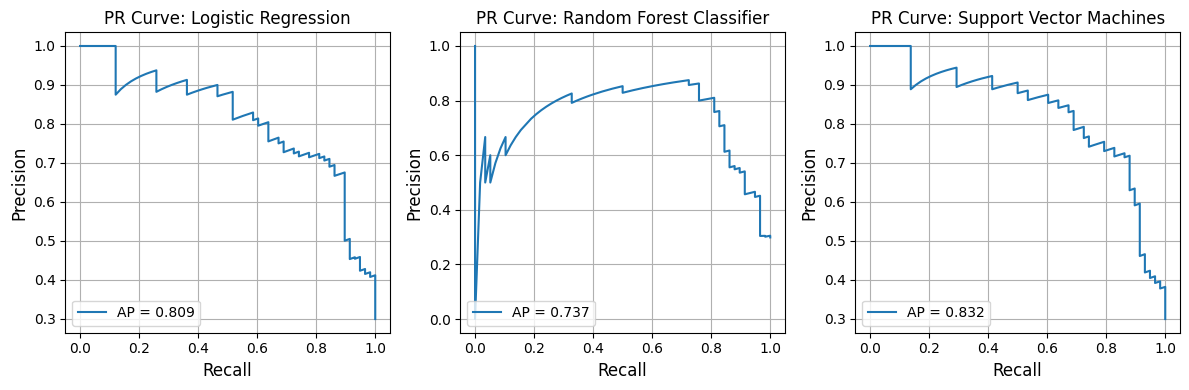
from model_metrics import show_pr_curve# Plot PR curves = pipelines_or_models,= X,= y,# x_label="Hello", = model_titles,= 3 ,= False ,= True ,= True ,= image_path_png,= image_path_svg,= (12 , 4 ),= 3 ,# tick_fontsize=16, # label_fontsize=16, # grid=True, # gridlines=False,
Running k-fold model metrics...
Processing Folds: 100%|██████████| 10/10 [00:00<00:00, 39.78it/s]
Average Precision for Logistic Regression: 0.809
Running k-fold model metrics...
Processing Folds: 100%|██████████| 10/10 [00:01<00:00, 5.19it/s]
Average Precision for Random Forest Classifier: 0.737
Running k-fold model metrics...
Processing Folds: 100%|██████████| 10/10 [00:00<00:00, 62.43it/s]
Average Precision for Support Vector Machines: 0.832
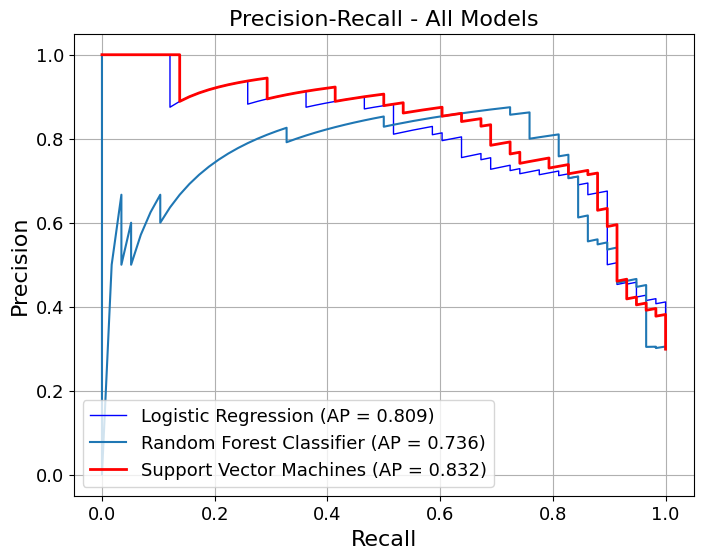
= pipelines_or_models,= X,= y,= True ,= model_titles,= "Precision-Recall - All Models" ,= {"Logistic Regression" : {"color" : "blue" , "linewidth" : 1 },"Random Forest" : {"color" : "lightblue" , "linewidth" : 1 },"Support Vector Machines" : {"color" : "red" ,"linestyle" : "-" ,"linewidth" : 2 ,= True ,= False ,= 3 ,= (8 , 6 ),# gridlines=False, = 16 ,= 13 ,= image_path_png,= image_path_svg,
Running k-fold model metrics...
Processing Folds: 100%|██████████| 10/10 [00:00<00:00, 40.17it/s]
Average Precision for Logistic Regression: 0.809
Running k-fold model metrics...
Processing Folds: 100%|██████████| 10/10 [00:02<00:00, 4.64it/s]
Average Precision for Random Forest Classifier: 0.736
Running k-fold model metrics...
Processing Folds: 100%|██████████| 10/10 [00:00<00:00, 42.88it/s]
Average Precision for Support Vector Machines: 0.832